
empty or not) checked for correlations among each other and with other features and in a second step for correlations with the label before a decision on ommitting them is made. Instead of simply dropping these columns, they are converted into binary features (i.e. Many parameters are available allowing a more restrictive data cleaning where needed.įurthermore, the function klib.mv_col_handling() provides a sophisticated selection mechanism for columns with relatively many missing values. Using this procedure, 56006 duplicate rows are identified in the subset, i.e., 56006 rows in 10 columns are encoded into a single column of dtype integer, greatly reducing the memory footprint and number of columns which should speed up model training.Īll of these functions were run with their relatively “soft” default settings. This allows us to pool and encode “carrier” and similar columns, while “tailnum” remains in the dataset. While this is unlikely, it is advised to specifically exclude features that provide sufficient informational content by themselves as well as the target column by using the “exclude” setting.Īs can be seen in *cat_plot()* the “carrier” column is made up of a few very frequent values - the top 4 values account for roughly 75% - while in “tailnum” the top 4 values barely make up 2%. To install this package run one of the following: conda install -c conda-forge klib. While the encoding itself does not lead to a loss in information, some details might get lost in the aggregation step. These are then added to the original data what allows dropping the previously identified and now encoded columns. Specifically, the pooling is achieved by finding duplicates in subsets of the data and encoding the largest possible subset with sufficient duplicates with integers. This function “pools” columns together based on several settings.  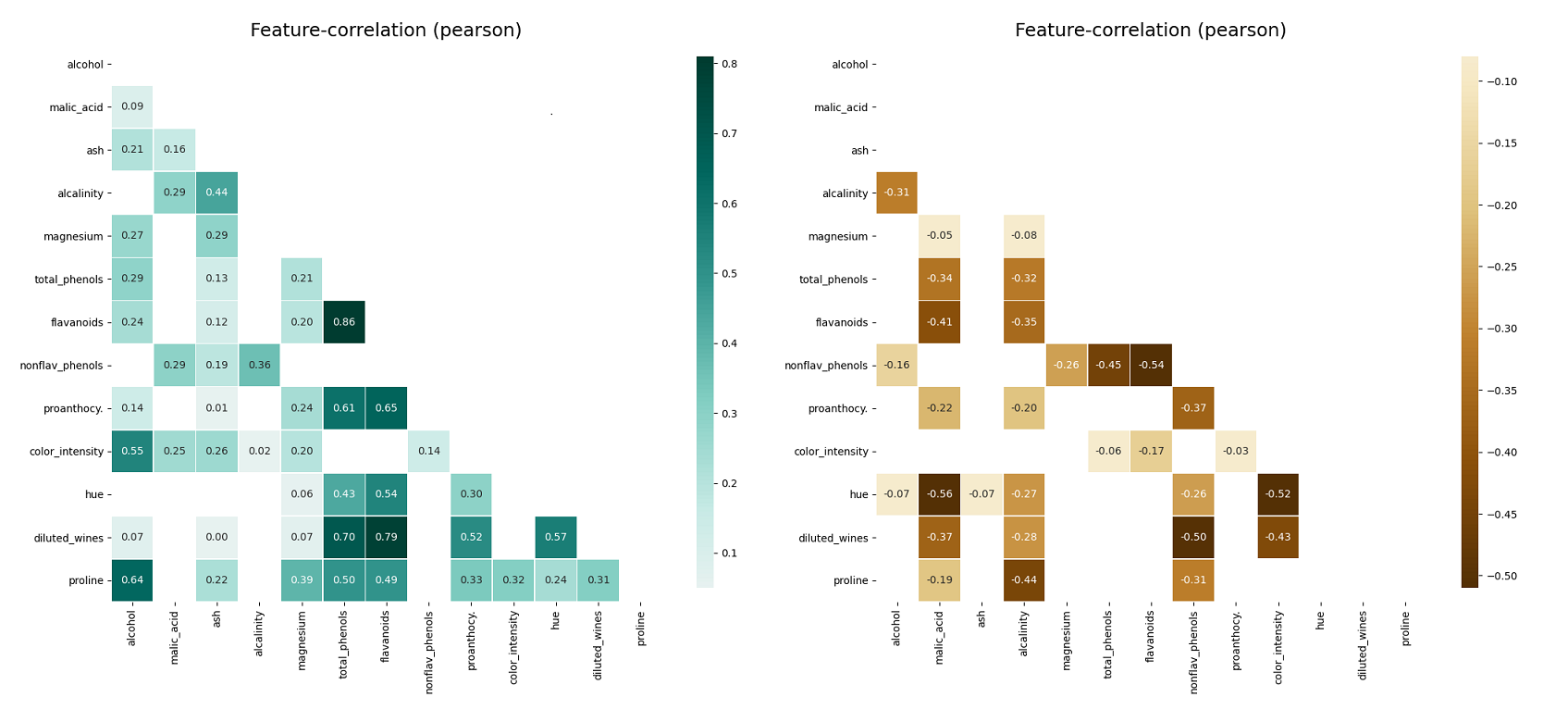
loss of information Examplesįind all available examples as well as applications of the functions in klib.clean() with detailed descriptions here.Further, klib.pool_duplicate_subsets() can be applied, what ultimately reduces the dataset to only 3.8 MB (from 51 MB originally). pool_duplicate_subsets( df) # pools subset of cols based on duplicates with min. mv_col_handling( df) # drops features with high ratio of missing vals based on informational content - klib. drop_missing( df) # drops missing values, also called in data_cleaning() - klib. convert_datatypes( df) # converts existing to more efficient dtypes, also called inside data_cleaning() - klib. clean_column_names( df) # cleans and standardizes column names, also called inside data_cleaning() - klib. data_cleaning( df) # performs datacleaning (drop duplicates & empty rows/cols, adjust dtypes.) - klib. missingval_plot( df) # returns a figure containing information about missing values # klib.clean - functions for cleaning datasets - klib. Based on project statistics from the GitHub repository for the PyPI package klib, we found that it has been starred 295 times, and that 0 other projects in the ecosystem are dependent on it. To begin, we must look for or create a data set. As such, we scored klib popularity level to be Limited. using Python, Pandas, Seaborn and Plotly Exploratory data analysis (EDA) refers to the process of performing investigation on one or many datasets to discover trends, spot anomalies, and test assumptions with the help of statistics and visualisations. dist_plot( df) # returns a distribution plot for every numeric feature - klib. The PyPI package klib receives a total of 842 downloads a week. corr_plot( df) # returns a color-encoded heatmap, ideal for correlations - klib. 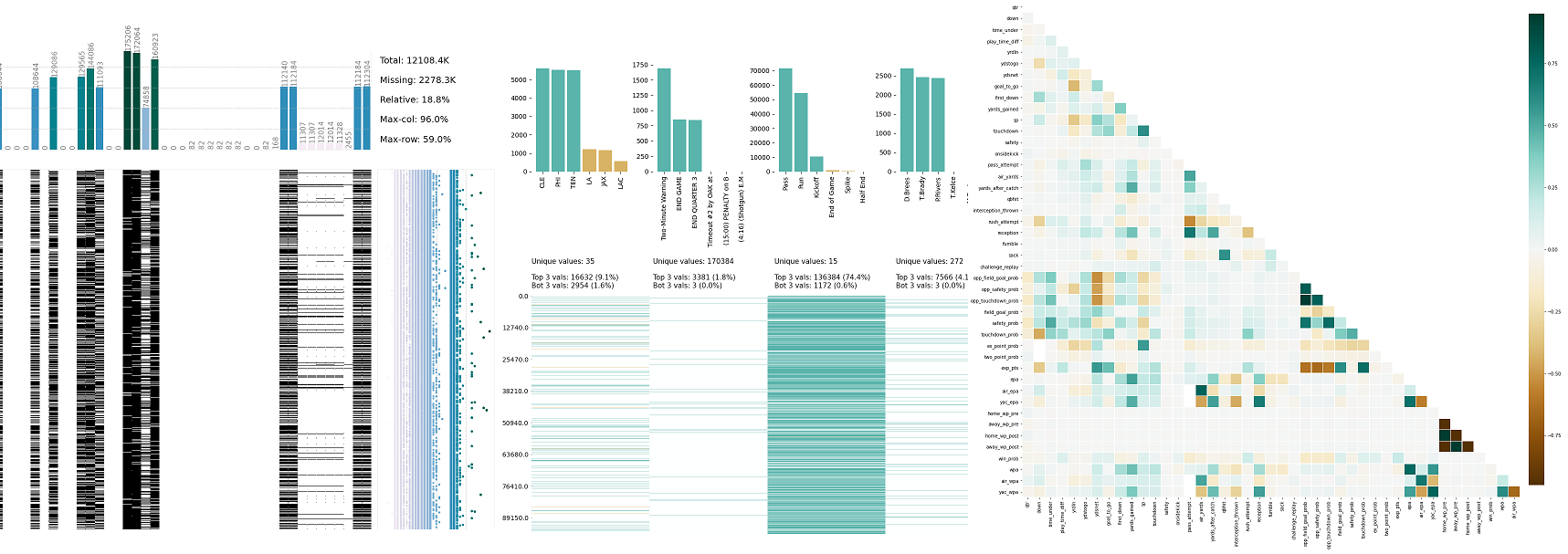
corr_mat( df) # returns a color-encoded correlation matrix - klib. cat_plot( df) # returns a visualization of the number and frequency of categorical features - klib. 
# scribe - functions for visualizing datasets - klib. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
